VBQs Class 12 Biology Biotechnology Principles and Processes
Very Short Answer Type Questions
Question. PCR requires very high temperature conditions where most of the enzymes get denatured. How was this problem resolved in a PCR ?
Answer : The thermostable DNA polymerase is used which is isolated from a bacterium called Thermus aquaticus.
It remains active during the high temperature this enzyme does not induce denaturation of DNA. This enzymes, taq polymerase carries out amplification of DNA at high temperature.
Question. Name the source of the DNA polymerase used in PCR technique. Mention why it is used ?
Answer : Thermus aquaticus, is the source of DNA polymerase because it is a heat-stable DNA polymerase.
Question. Write the function of a bioreactor.
Answer : Bioreactors are required to produce large volumes (100 – 1000 litres) of recombinant proteins / desired protein / enzymes.
Short Answer Type Questions
Question. (i) A recombinant vector with a gene of interest inserted within the gene of β-galactosidase enzyme, is introduced into a bacterium. Explain the method that would help in selection of recombinant colonies from non-recombinant ones.
(ii) Why is this method of selection referred to as “insertional inactivation” ?
Answer : (i) Bacteria are grown in a medium with chromogenic substrate, colonies formed show blue colour-no recombinants, no blue colour –presence of recombinants.
(ii) Gene for the enzyme is inactivated by insertion.
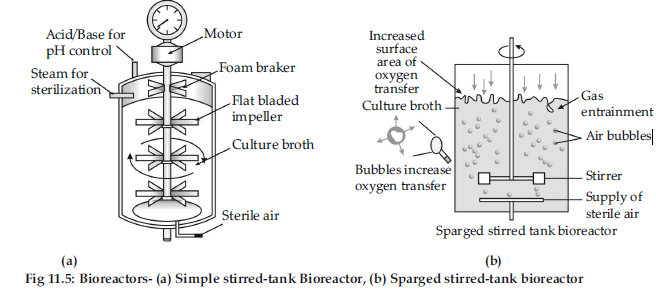
Question. Name two commonly used bioreactors. State the importance of using bioreactors.
Answer : The two types of commonly used bioreactors are:
(i) Stirred tank bioreactor and
(ii) Sparged stirred tank bioreactors.
Importance :
A bioreactor also called fermenter is a specialized container or a vessel required for the production of a number of industrial products such as antibiotics, enzymes and vitamins with the help of microorganisms and microbial reactions going on inside the fermenter.
In the bioreactor, a huge amount of raw material or substrates are biologically converted into specific industrial products like vitamins, enzymes, antibiotics, etc. on the industrial scale.
Question. How can bacterial DNA be released from the bacterial cell for biotechnology experiments ?
Answer : DNA is enclosed within the membranes; so, we have to break and open the cell to release DNA along with other macromolecules such as RNA, proteins, polysaccharides and also lipids. This can be achieved by treating the bacterial cell / plant or animal tissue with enzymes such as lysozyme (bacteria), cellulase (plant cell), chitinase (fungus).
Question. Explain three steps involved in polymerase chain reaction.
OR
Suggest and describe a technique to obtain multiple copies of a gene of interest in vitro.
OR
Any recombinant DNA with a desired gene is required in billion copies for commercial use.
How is the amplification done ? Explain.
OR
How is the amplification of a gene sample of interest carried out using Polymerase Chain Reaction (PCR) ?
Answer : (i) Denaturation: Two strands of DNA are separated by heating.
(ii) Annealing : One set of primers are attached / annealed to the separated DNA (reverse and forward primer) strands.
(iii) Extension: Taq polymerase catalyses the extension of primers using genomic DNA as template and nucleotides provided in the reaction.
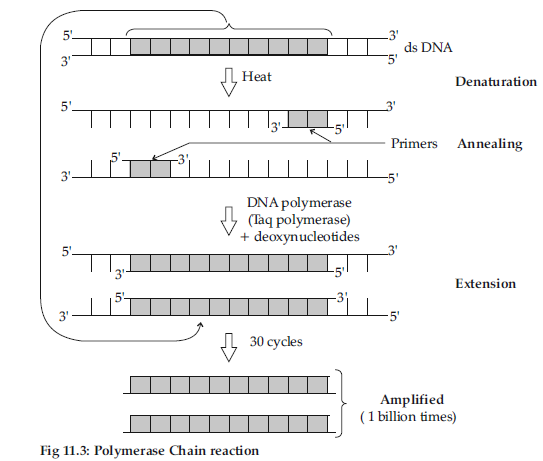
Detailed Answer :
Polymerase Chain Reaction Technique can be used to obtain multiple copies of a gene of interest in vitro.
The PCR consists of three steps :
(i) Denaturation : The segment of double stranded DNA of interest is heated to separate the two strands at high temperature of about 90-95°C.
(ii) Annealing : A set of primers (chemically synthesised oligonucleotides which are complementary to the regions of DNA) are annealed to both the separated DNA segments.
DNA polymerase (Taq polymerase) extends the primer in the 5’ to 3’ direction.
(iii) Extension : The separated DNA segments acts as templates and primers synthesises new strands along the entire length of DNA strands.
(iv) Amplification : The cycle is repeated several times to generate up to one billion identical copies of DNA.
Question. (i) Why was a bacterium used in the first instance of the construction of an artificial recombinant DNA molecule?
(ii) Name the scientists who accomplished this and how?
Answer : (i) Bacterium has a plasmid in which the desired gene is introduced / the gene to be transferred is from a bacterium. The anti-biotic resistance gene / the host cell (bacterial cell) is required for gene cloning / bacteria produce restriction endonucleases.
(ii) Herbert Boyer and Stanley Cohen Antibiotic resistant gene was isolated using restriction enzyme and introduced into the plasmid of bacterium Salmonella typhimurium.
Later the recombinant plasmid was introduced into the bacterium E. coli. so that it could make copies of gene.
Question. Draw a labelled sketch of sparged – stirred-tank bioreactor. Write its application.
Answer : Application – Bioreactors are used to produce larger biomass leading to higher yields of desired protein /recombinant protein in bioreactors. Thus, carries the processing of large volume of culture / conversion of raw materials into specific product biologically.
Question. (i) List the three steps involved in Polymerase Chain Reaction (PCR).
(ii) Name the source organism of Taq polymerase.
Explain the specific role of this enyme in PCR.
Answer : (i) (a) Denaturation (b) Annealing (c) Extension.
(ii) Thermus aquaticus, it remains active during the high temperature, (induced to denature double stranded DNA) and catalyses polymerisation of DNA.
Question. Describe the roles of heat, primers and the bacterium Thermus aquaticus in the process of PCR.
Answer : Heat : Denaturation / separation of DNA into two strands.
Primer : Enzyme DNA Polymerase extend the primers using the nucleotides provided in the reaction and the genomic DNA as template.
Thermus aquatics : Source of thermostable DNA polymerase i.e., Taq polymerase.
Question. How does β-galactosidase coding sequence act as a selectable marker ? Explain. Why it is a preferred selectable marker to antibiotic resistant genes ?
Answer : (i) Presence of a chromogenic substrate gives blue colour, if the plasmid in the bacteria does not have an insert (non-recombinants).
(ii) With the insert-do not produce any colour, recombinant colonies.
(iii) Selection of recombinants due to inactivation of antibiotics, requires simultaneous plating on two plates having different antibiotics / process is more cumbersome.
Question. Explain three basic steps to be followed during genetic modification of an organism.
Answer : (i) Identification of DNA with desirable genes, so that the genetically modified organism has largely desirable genes.
(ii) Introduction of the DNA with desirable genes, into the host using vector.
(iii) Maintenance of introduced DNA in the host and transfer of the DNA to its progeny through cloning.
Question. A schematic representation of polymerase chain reaction (PCR) upto extension stage is given below.
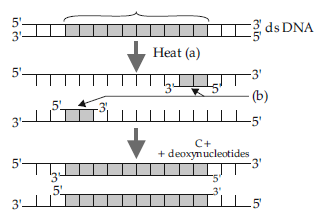
Answer the questions that follows :
(i) Name the process ‘a’
(ii) Identify ‘b’
(iii) Identify ‘c’ and mention its importance in PCR.
Answer : (i) Denaturation process.
(ii) Primers.
(iii) Taq DNA polymerase.
Taq polymerase is a thermostable enzyme isolated from a thermophilic bacterium Thermus aquaticus. This enzyme remains functional during high temperature required for extension of DNA.
Long Answer Type Questions
Question. (i) Describe the different steps in one complete cycle of PCR.
(ii) State the purpose of such an amplified DNA sequence.
Answer : (i) The presence of chromogenic substrate gives blue coloured colonies, in presence of β-galactosidase.
Presence of an insert (recombinant DNA) results into inactivation of the enzyme, colonies with inactivation of β-galactosidase do not produce any colour.
Detailed Answer :
The insertion of rDNA into the coding sequence of an enzyme alpha-galactosidase leads to the inactivation of the enzyme called insertional inactivation. The recombinants do not produce a blue coloured colonies in the presence of chromogenic substrate while the non-recombinants produce a blue colour.
(ii) Purpose : Used to ligate with a vector for further cloning. Detection of bacteria or virus by amplification of their DNA / detection of HIV in AIDS patient. Detection of mutation in genes in suspected cancer patients.
